Menu
Conclusions
| dumar_2-21-17_rough_draft_final_talk.pdf | |
| File Size: | 332 kb |
| File Type: | |
| dumar_4-4-17_rough_draft_final_talk.pdf | |
| File Size: | 996 kb |
| File Type: | |
| dumar_final_talk_4-18-17.pdf | |
| File Size: | 1526 kb |
| File Type: | |
Introduction
|
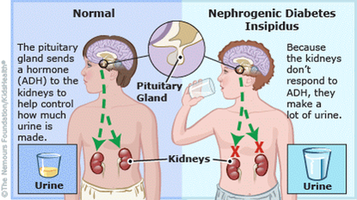
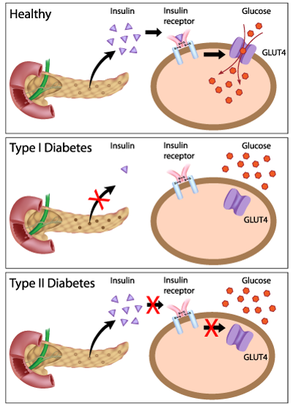
Figure 1: Comparison of the physiology leading to NDI(left) versus diabetes mellitus(right).
|
Nephrogenic diabetes insipidus(NDI) is a disorder of improper water balance within the kidney that results in symptoms of excessive thirst and excessive urine production (1). NDI is similar to the more widely known diabetes mellitus, in that both diseases share the symptom of excessive urine production(2). However, NDI is caused by dysfunction in how the kidney interacts with anti-diuretic hormone, while diabetes mellitus is caused by dysfunction of the body's response to insulin(3,4, Figure 1).
|
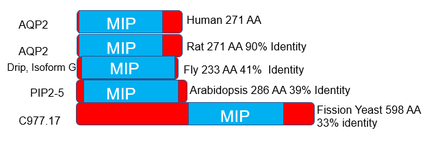
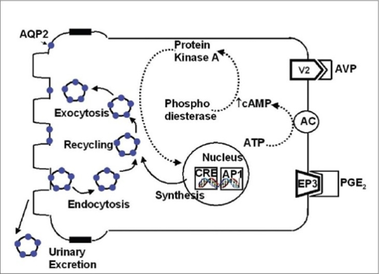
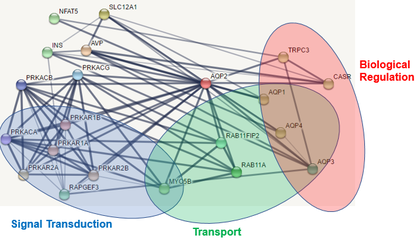
There are multiple variants of NDI, but the variants that are autosomally inherited are caused by defects in the aquaporin-2(AQP2) gene (5). AQP2 is a water transport protein associated with the plasma membrane of kidney cells, and its levels at the cell surface are regulated by vesicular transport (6,Figure 2). AQP2 is well conserved across a wide variety of organisms due to the importance of water transport to all organisms(Figure 3). AQP2 interacts with a variety of other proteins, particularly those involved in biological regulation, transport, and signal transduction(Figure 4). Phosphorylation of the C-terminus of AQP2 is important for this vesicular transport, but the role phosphorylation events play at all residues is not well understood(7). In particular, serine 264(S264) has not been thoroughly investigated, other than prior studies noting that AQP2 with phosphorylation at S264 showed reduced endocytosis (8).
I hypothesize that phosphorylation of S264 is used to promote retention of AQP2 at the plasma membrane.
This will be tested at three levels of complexity: at position 264 within the AQP2 protein, at the phosphorylation state of position 264, and at the level of protein interactors of AQP2 when position 264 is in different phsophorylation states. Mice will be used as model organisms for these studies, as they are easily manipulated genetically and the signaling pathways involved in AQP2 movement to the plasma membrane are conserved between mice and humans(9).
I hypothesize that phosphorylation of S264 is used to promote retention of AQP2 at the plasma membrane.
This will be tested at three levels of complexity: at position 264 within the AQP2 protein, at the phosphorylation state of position 264, and at the level of protein interactors of AQP2 when position 264 is in different phsophorylation states. Mice will be used as model organisms for these studies, as they are easily manipulated genetically and the signaling pathways involved in AQP2 movement to the plasma membrane are conserved between mice and humans(9).
Figure 2: Cycling of AQP2 within the cell
Specific Aim 1
|
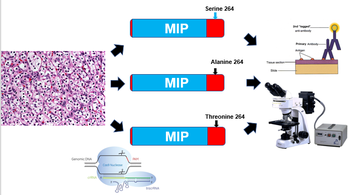
Figure 5: Transformation of mouse cell lines using CRISPR-Cas9 and evaluation of AQP2 location using fluorescent antibodies
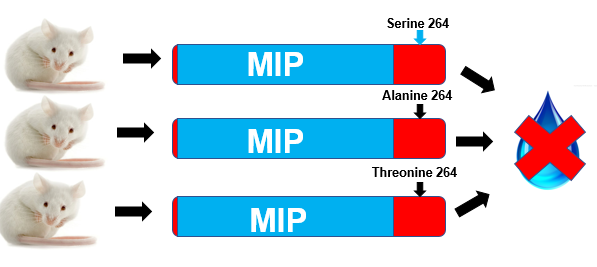
Figure 6: Water deprivation tests on mice that have been transformed using CRISPR-Cas9.
|
In the first aim, natural populations of mice will be sequenced for variants of AQP2 that are variable at position 264 using an Illumnia Mi-seq(10). Once natural variants at position 264 have been identified, these mutations will be engineered into mouse kidney cell lines using CRISPR-Cas9 technology(11). The localization of AQP2 within these cell lines will then be tracked using fluorescent antibodies specific to AQP2(12,Figure 5)Additionally, the same AQP2 mutations will be engineered into mice using CRISPR-Cas9 technology(13). The urine output of these mice will be evaluated using a water deprivation test, with special consideration to avoid acute water stress in the mice (14,Figure 6). Water deprivation tests are used in humans to diagnose NDI, and will thus also serve as a good phenotypic assay for NDI symptoms in the mice(15).
I hypothesize that mutations that all mutations that affect S264 will lead to observable phenotypes in cellular and organismal assays. These analyses will determine if the amino acid at site 264 in AQP2 is able to alter the location of AQP2 within mouse cells or alter urinary output of the mouse. Alterations in urinary output or AQP2 localization would provide support for previous observations of the importance of S264 in AQP2 trafficking (8). |
Specific Aim 2
|
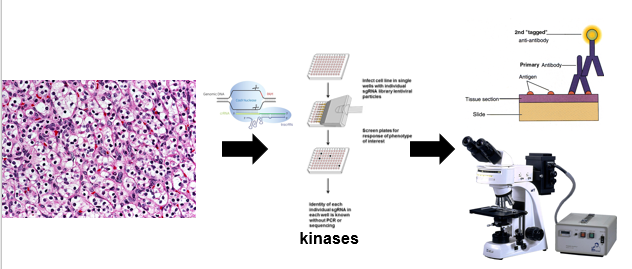
Figure 7: CRISPR screen of kinases in mouse cell lines.
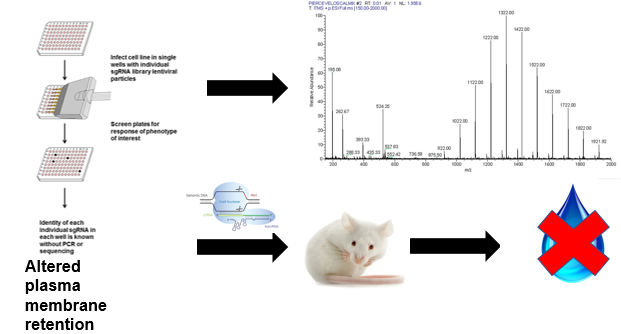
Figure 8: Mass spectrometry and water deprivation test evaluation of kinase deletions.
|
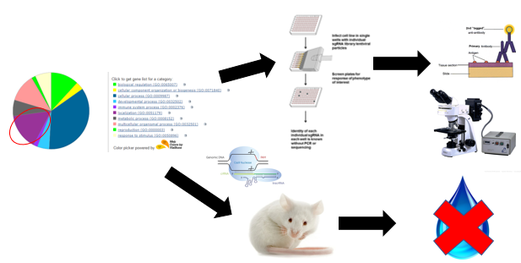
In the second aim, the importance of the phosphorylation state of S264 for AQP2 trafficking will be evaluated using a CRISPR-Cas9 screen(16). Kinases are enzymes that phsophorylate a wide variety of proteins, and they are extremely important in signaling pathways(17). CRISPR screening will be used on mouse kidney cell lines to generate cell populations with particular kinases deleted, and AQP2 location within the cell lines will be evaluated using fluorescent antibodies as in the first aim.Those cell lines showing altered movement of AQP2 will then be further analyzed using mass spectrometry to determine the phosphorylation state of AQP2 within these cells(18). Additionally, mice will be generated with the same kinase deletions as those analyzed using mass spectrometry and water deprivation tests will be done in those mice.
I hypothesize that kinase deletions that affect the phosphorylation state at S264 within AQP2 will lead to abnormalities in AQP2 trafficking and in water deprivation tests. These analyses will provide insight into if the phosphorylation state of S264 is responsible for its effects on AQP2 location within the cell. |
Specific Aim 3
|
Figure 9: Altered AQP2 constructs that simulate permanent phoshporylation or inability to be phosphorylated.
Figure 10: Cellular and organismal analysis of deletions of localization involved interactors with AQP2 that are changed based on site 264 phosphorylation state.
|
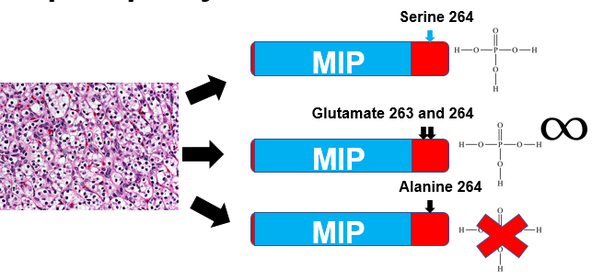
In the third aim, the effect of altered phosphorylation state at S264 in AQP2 on protein interactions will be analyzed. Two different AQP2 constructs will be used for this experiment, one in which S264 is replaced by an alanine and one in which S264 is replaced by two adjacent glutamine residues. Alanine residues cannot be phosphorylated and thus simulate a permanently unphosphorylated state of S264, while two adjacent glutamine residues have side chains that simulate the bulk and negative charge of a phosphate(19). These AQP2 constructs will be engineered into cell lines using CRISPR-Cas9, and then co-immunoprecipitation will be used to evaluate interactions of altered AQP2 constructs(20).The ontology of proteins whose interactions are altered by changes in phosphorylation at the 264 site in AQP2 will be evaluated, and those involved in localization within the cell will be further analyzed. CRISPR-Cas9 will be used to create mice and cell lines that have deletions of localization involved interactors, and AQP2 localization and urine output of mice will be evaluated for each interactor.
I hypothesize that alanine substitued AQP2 will interact with more proteins associated with endocytosis than either of the other two AQP2 constructs. This analysis will determine if the effects of phosphorylation state at site 264 in AQP2 are due to differences in protein regulators. |
Future Directions
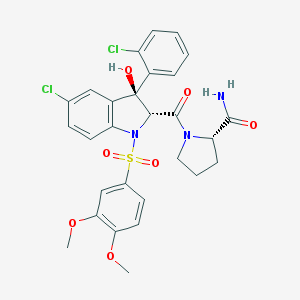
There are currently drugs available to treat NDI caused by dysfunctions in the receptor for ADH, but not for NDI caused by AQP2 dysfunction(21). Using the differently regulated protein interactors found within the third aim of this study, a possible future avenue of research would be the development of novel therapeutics that target one of these protein interactors(22). These drugs could then be used to lessen the impact that NDI has on people's daily lives.
Figure 11: Structure of relcovapatan, a compound used
to treat X-linked NDI.
to treat X-linked NDI.
1. Nephrogenic Diabetes Insipidus. National Organization for Rare Disorders. 2016. https://rarediseases.org/rare-diseases/nephrogenic-diabetes-insipidus/. Accessed May 11th, 2017.
2.Description. Nephrogenic Diabetes Insipidus Foundation. http://www.ndif.org/pages/4-Description Accessed May 11th,2017
3. Diabetes Insipidus. The Pituitary Foundation 2017. https://www.pituitary.org.uk/information/pituitary-conditions/diabetes-insipidus/. Accessed May 11th,2017.
4. Introduction to diabetes mellitus. Frits Holleman. https://www.diapedia.org/introduction-to-diabetes-mellitus/1104085113. Accessed May 11th,2017.
5.Vanlieburg, A. F., Verdijk, M. A. J., Knoers, V., Vanessen, A. J., Proesmans, W., Mallmann, R., . . . Deen, P. M. T. (1994). Patients with autosomal nephrogenic diabetes-insipidus homozygous for mutations in the aquaporin-2 water-channel gene. American Journal of Human Genetics, 55(4), 648-652.
6.Mulders, S. M., Bichet, D. G., Rijss, J. P. L., Kamsteeg, E. J., Arthus, M. F., Lonergan, M., . . . Deen, P. M. T. (1998). An aquaporin-2 water channel mutant which causes autosomal dominant nephrogenic diabetes insipidus is retained in the Golgi complex. Journal of Clinical Investigation, 102(1), 57-66.
7.Jung, H. J., & Kwon, T. H. (2016). Molecular mechanisms regulating aquaporin-2 in kidney collecting duct. American Journal of Physiology-Renal Physiology, 311(6), F1318-F1328.
8.Ren, H. W., Yang, B. X., Ruiz, J. A., Efe, O., Ilori, T. O., Sands, J. M., & Klein, J. D. (2016). Phosphatase inhibition increases AQP2 accumulation in the rat IMCD apical plasma membrane. American Journal of Physiology-Renal Physiology, 311(6), F1189-F1197.
9.Ando, F., Sohara, E., Morimoto, T., Yui, N., Nomura, N., Kikuchi, E., . . . Uchida, S. (2016). Wnt5a induces renal AQP2 expression by activating calcineurin signalling pathway. Nature Communications, 7.
10.Key Applications and Methods. Illumina 2017. https://www.illumina.com/systems/sequencing-platforms/miseq/applications.html
11.Yang, M. Z., Zhang, L., Stevens, J., & Gibson, G. (2014). CRISPR/Cas9 mediated generation of stable chondrocyte cell lines with targeted gene knockouts; analysis of an aggrecan knockout cell line. Bone, 69, 118-125.
12.Tokuda, M., Fujisawa, M., Miyashita, K., Kawakami, Y., Morimoto-Yamashita, Y., & Torii, M. (2015). Involvement of TRPV1 and AQP2 in hypertonic stress by xylitol in odontoblast cells. Connective Tissue Research, 56(1), 44-49.
13.Qin, W. N., Dion, S. L., Kutny, P. M., Zhang, Y. F., Cheng, A. W., Jillette, N. L., . . . Wang, H. Y. (2015). Efficient CRISPR/Cas9-Mediated Genome Editing in Mice by Zygote Electroporation of Nuclease. Genetics, 200(2), 423-+.
14.Bekkevold, C. M., Robertson, K. L., Reinhard, M. K., Battles, A. H., & Rowland, N. E. (2013). Dehydration Parameters and Standards for Laboratory Mice. Journal of the American Association for Laboratory Animal Science, 52(3), 233-239.
15.Sahakitrungruang, T., Wacharasindhu, S., Sinthuwiwat, T., Supornsilchai, V., Suphapeetiporn, K., & Shotelersuk, V. (2008). Identification of two novel aquaporin-2 mutations in a Thai girl with congenital nephrogenic diabetes insipidus. Endocrine, 33(2), 210-214.
16.Song, C. Q., Li, Y. X., Mou, H. W., Moore, J., Park, A., Pomyen, Y., . . . Xue, W. (2017). Genome-Wide CRISPR Screen Identifies Regulators of Mitogen-Activated Protein Kinase as Suppressors of Liver Tumors in Mice. Gastroenterology, 152(5), 1161-+.
17.Nishizuka, Y. (1984). Protein-kinases in signal transduction. Trends in Biochemical Sciences, 9(4), 163-166.
18.Dephoure, N., Gould, K. L., Gygi, S. P., & Kellogg, D. R. (2013). Mapping and analysis of phosphorylation sites: a quick guide for cell biologists. Molecular Biology of the Cell, 24(5), 535-542.
19.Hunter, T. (2012). Why nature chose phosphate to modify proteins. Philosophical Transactions of the Royal Society B-Biological Sciences, 367(1602), 2513-2516.
20.Moeller, H. B., Slengerik-Hansen, J., Aroankins, T., Assentoft, M., MacAulay, N., Moestrup, S. K., . . . Fenton, R. A. (2016). Regulation of the Water Channel Aquaporin-2 via 14-3-3 theta and-zeta. Journal of Biological Chemistry, 291(5), 2469-2484
21.Gavrin, L. K., Denny, R. A., & Saiah, E. (2012). Small Molecules That Target Protein Misfolding. Journal of Medicinal Chemistry, 55(24), 10823-10843.
22. Stockwell, B. R. (2004). Exploring biology with small organic molecules. Nature, 432(7019), 846-854.
2.Description. Nephrogenic Diabetes Insipidus Foundation. http://www.ndif.org/pages/4-Description Accessed May 11th,2017
3. Diabetes Insipidus. The Pituitary Foundation 2017. https://www.pituitary.org.uk/information/pituitary-conditions/diabetes-insipidus/. Accessed May 11th,2017.
4. Introduction to diabetes mellitus. Frits Holleman. https://www.diapedia.org/introduction-to-diabetes-mellitus/1104085113. Accessed May 11th,2017.
5.Vanlieburg, A. F., Verdijk, M. A. J., Knoers, V., Vanessen, A. J., Proesmans, W., Mallmann, R., . . . Deen, P. M. T. (1994). Patients with autosomal nephrogenic diabetes-insipidus homozygous for mutations in the aquaporin-2 water-channel gene. American Journal of Human Genetics, 55(4), 648-652.
6.Mulders, S. M., Bichet, D. G., Rijss, J. P. L., Kamsteeg, E. J., Arthus, M. F., Lonergan, M., . . . Deen, P. M. T. (1998). An aquaporin-2 water channel mutant which causes autosomal dominant nephrogenic diabetes insipidus is retained in the Golgi complex. Journal of Clinical Investigation, 102(1), 57-66.
7.Jung, H. J., & Kwon, T. H. (2016). Molecular mechanisms regulating aquaporin-2 in kidney collecting duct. American Journal of Physiology-Renal Physiology, 311(6), F1318-F1328.
8.Ren, H. W., Yang, B. X., Ruiz, J. A., Efe, O., Ilori, T. O., Sands, J. M., & Klein, J. D. (2016). Phosphatase inhibition increases AQP2 accumulation in the rat IMCD apical plasma membrane. American Journal of Physiology-Renal Physiology, 311(6), F1189-F1197.
9.Ando, F., Sohara, E., Morimoto, T., Yui, N., Nomura, N., Kikuchi, E., . . . Uchida, S. (2016). Wnt5a induces renal AQP2 expression by activating calcineurin signalling pathway. Nature Communications, 7.
10.Key Applications and Methods. Illumina 2017. https://www.illumina.com/systems/sequencing-platforms/miseq/applications.html
11.Yang, M. Z., Zhang, L., Stevens, J., & Gibson, G. (2014). CRISPR/Cas9 mediated generation of stable chondrocyte cell lines with targeted gene knockouts; analysis of an aggrecan knockout cell line. Bone, 69, 118-125.
12.Tokuda, M., Fujisawa, M., Miyashita, K., Kawakami, Y., Morimoto-Yamashita, Y., & Torii, M. (2015). Involvement of TRPV1 and AQP2 in hypertonic stress by xylitol in odontoblast cells. Connective Tissue Research, 56(1), 44-49.
13.Qin, W. N., Dion, S. L., Kutny, P. M., Zhang, Y. F., Cheng, A. W., Jillette, N. L., . . . Wang, H. Y. (2015). Efficient CRISPR/Cas9-Mediated Genome Editing in Mice by Zygote Electroporation of Nuclease. Genetics, 200(2), 423-+.
14.Bekkevold, C. M., Robertson, K. L., Reinhard, M. K., Battles, A. H., & Rowland, N. E. (2013). Dehydration Parameters and Standards for Laboratory Mice. Journal of the American Association for Laboratory Animal Science, 52(3), 233-239.
15.Sahakitrungruang, T., Wacharasindhu, S., Sinthuwiwat, T., Supornsilchai, V., Suphapeetiporn, K., & Shotelersuk, V. (2008). Identification of two novel aquaporin-2 mutations in a Thai girl with congenital nephrogenic diabetes insipidus. Endocrine, 33(2), 210-214.
16.Song, C. Q., Li, Y. X., Mou, H. W., Moore, J., Park, A., Pomyen, Y., . . . Xue, W. (2017). Genome-Wide CRISPR Screen Identifies Regulators of Mitogen-Activated Protein Kinase as Suppressors of Liver Tumors in Mice. Gastroenterology, 152(5), 1161-+.
17.Nishizuka, Y. (1984). Protein-kinases in signal transduction. Trends in Biochemical Sciences, 9(4), 163-166.
18.Dephoure, N., Gould, K. L., Gygi, S. P., & Kellogg, D. R. (2013). Mapping and analysis of phosphorylation sites: a quick guide for cell biologists. Molecular Biology of the Cell, 24(5), 535-542.
19.Hunter, T. (2012). Why nature chose phosphate to modify proteins. Philosophical Transactions of the Royal Society B-Biological Sciences, 367(1602), 2513-2516.
20.Moeller, H. B., Slengerik-Hansen, J., Aroankins, T., Assentoft, M., MacAulay, N., Moestrup, S. K., . . . Fenton, R. A. (2016). Regulation of the Water Channel Aquaporin-2 via 14-3-3 theta and-zeta. Journal of Biological Chemistry, 291(5), 2469-2484
21.Gavrin, L. K., Denny, R. A., & Saiah, E. (2012). Small Molecules That Target Protein Misfolding. Journal of Medicinal Chemistry, 55(24), 10823-10843.
22. Stockwell, B. R. (2004). Exploring biology with small organic molecules. Nature, 432(7019), 846-854.
Image References
•AQP2 transport image: https://openi.nlm.nih.gov/detailedresult.php?img=PMC2813137_IJN-18-95-g002&req=4
•Mass spectrum image: https://www.thermofisher.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/protein-biology-application-notes/mass-spec-calibration-solutions.html
•Mouse image: http://www.npr.org/templates/story/story.php?storyId=130653692
•Kidney cells image: https://directorsblog.nih.gov/2013/07/09/a-diabolical-recipe-for-aggressive-kidney-cancer/
•CRISPR/Cas9 image: http://dharmacon.gelifesciences.com/gene-editing/crispr-cas9/crispr-guide-rna/
•Antibody image: http://www.doctorc.net/Labs/Lab2/Examples/exfluor.htm
•Fluorescence microscope image: http://www.microscope.com/specialty-microscopes/epi-fluoresecence-microscopes/meiji-techno-mt6000-epi-fluorescence-microscope.html#gref
•Giant red X: http://www.clipartkid.com/red-x-cliparts/
•Water Droplet: http://clipartall.com/clipart/5046-water-droplet-clipart.html
•CRISPR screen image: http://www.genecopoeia.com/resource/genome-editing-applications-for-genecopoeia-crispr-sgrna-libraries/
•Phosphate molecule: http://web.chem.ucla.edu/~harding/IGOC/P/phosphate_group.html
•Infinity sign: http://www.deke.com/content/dekes-techniques-104-crafting-infinity-symbol-match-a-font-illustrator
•Co-immunoprecipitation: http://www.genscript.com/immunoprecipitation.html