Menu
Protein Interactions
This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What are Protein Interaction Networks?
Protein interaction networks are used to show the wide variety of additional proteins that a protein of interest interacts with. There are several different ways in which the data to support these interactions are determined, including yeast 2-hybrid assays and co-immunoprecipitation (1,2). By studying these interaction networks, new insights can be made into drug targets for therapeutics and the overall pathways that the protein is involved in can be evaluated (1).
Human AQP2 interactions
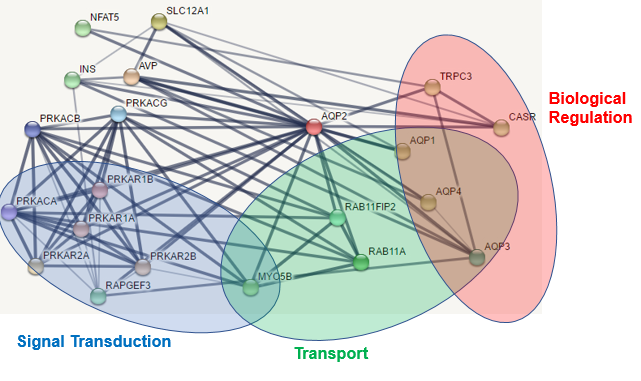
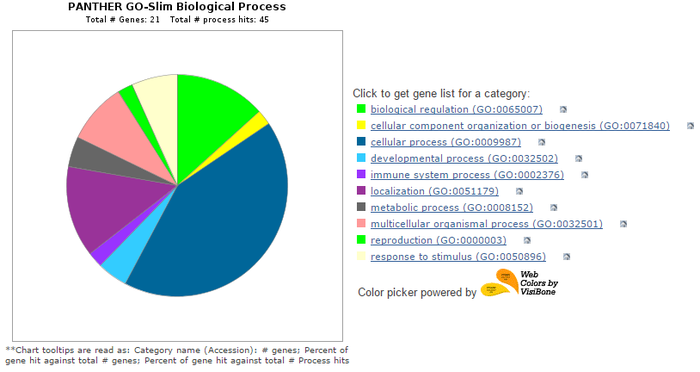
The figure at the left displays the protein interaction network for human AQP2(red dot in center) as generated using STRING (4). Below is a pie chart depicting the biological processes that the proteins identified in this network are involved in. The ontologies of these interactors were visualized using PANTHER (5). The most dominant functional categories of the interacting proteins are involvement in cellular processes(including signal transduction), localization(including transport), multicellular organism processes, and biological regulation. One result of note is the inclusion of several other aquaporins within this network, including AQP1, AQP3, and AQP4. This opens the possibility for some overlap in coregulation of these different aquaporins.
Mouse AQP2 interactions
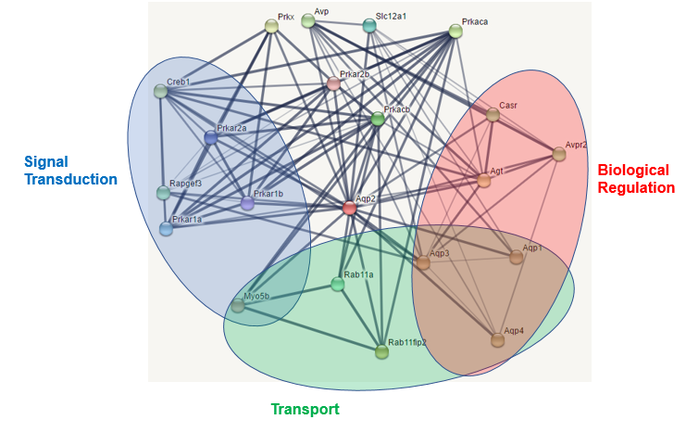
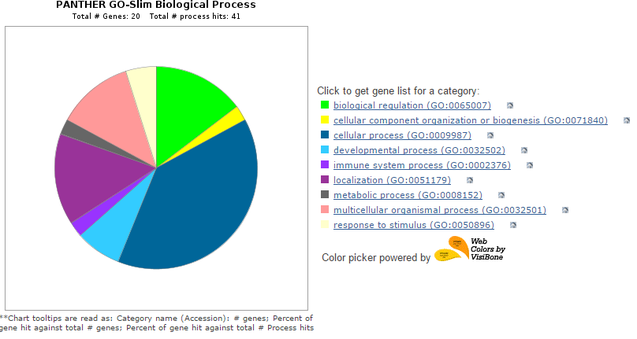
The image at left depicts the protein interaction network of mouse AQP2(red dot in center) as generated by STRING (4). As with the human network, a chart of the biological process gene ontologies of the interactors was generated using PANTHER (5). Like in humans, cellular process and biological regulation were highly represented. However, there is no reproduction annotation and a slight expansion of the multicellular organism process category. Despite this, three different aquaporin genes can still be found in this interaction network: AQP1, AQP3, and AQP4.
Discussion
The high degree of similarity between the biological processes associated with proteins interacting with mouse and human AQP2 along with conservation of many of the interacting proteins supports the use of mice as a model system for testing the processes controlling regulation of AQP2 at the plasma membrane within the kidney collecting duct. However, divergence of the functions of several genes can be seen between the two organsism, such as PRKACA in humans being associated with signal transduction while it is not in mice. These differences limit the overall scope of conclusions that can be made extrapolated from observed protein interactions in mice.
Works Cited
1.Hamdi, A., & Colas, P. (2012). Yeast two-hybrid methods and their applications in drug discovery. Trends in Pharmacological Sciences, 33(2), 109-118. doi:10.1016/j.tips.2011.10.008
2.Kuzmanov, U., & Emili, A. (2013). Protein-protein interaction networks: probing disease mechanisms using model systems. Genome Medicine, 5. doi:10.1186/gm441
3.STRING. http://string-db.org/. Accessed April 11, 2017.
4.PANTHER CLASSIFICATION SYSTEM. http://www.pantherdb.org/. Accessed April 11, 2017.
2.Kuzmanov, U., & Emili, A. (2013). Protein-protein interaction networks: probing disease mechanisms using model systems. Genome Medicine, 5. doi:10.1186/gm441
3.STRING. http://string-db.org/. Accessed April 11, 2017.
4.PANTHER CLASSIFICATION SYSTEM. http://www.pantherdb.org/. Accessed April 11, 2017.