Menu
AQP2 Domains
This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What are protein domains and how do you find them?
Domains are regions of a protein that have a conserved function across many different proteins (1). Analysis of protein structure to determine domains can be done using several programs, including PFAM(2) and SMART(3).
Domains of AQP2
AQP2 only showed evidence of a single MIP domain.
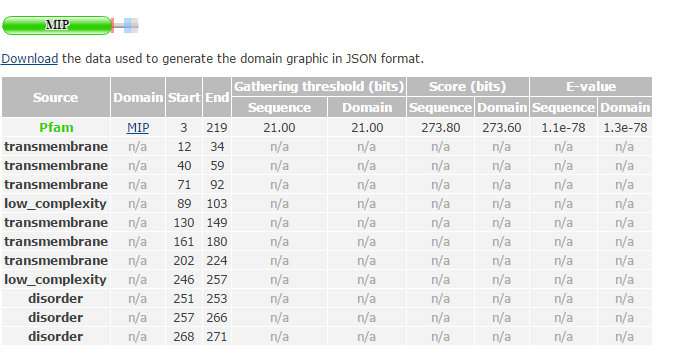
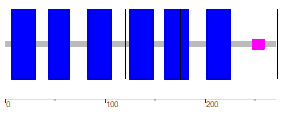
Figure 1: Domains of human AQP2 detected by PFAM. In addition to functional domains, PFAM also notates transmembrane regions, regions of low complexity, and intrinsically disordered regions(have little structure). The only domain that PFAM detected within AQP2 was MIP, also known as major intrinsic protein.
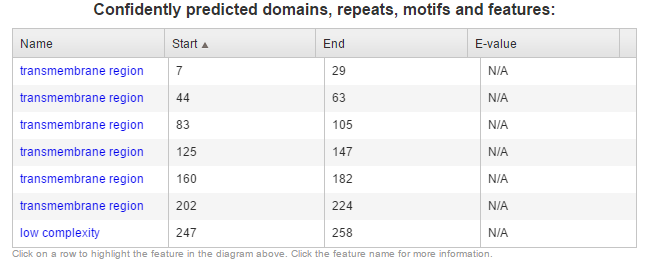
Figure 2: Domains of human AQP2 detected by SMART. SMART did not detect the MIP domain or intrinsically disordered regions. Locations of transmembrane and low complexity regions also differ slightly from PFAM's annotations.
How well conserved is the domain architecture of AQP2 and its homologues?
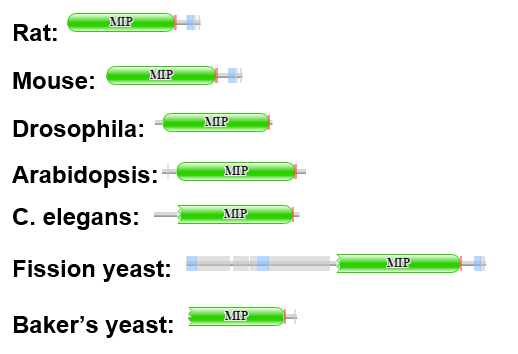
Figure 3: PFAM representation of AQP2 homologs. All of the homologs share the MIP domain and none of the homologs contain any additional domains beyond MIP.
Discussion of AQP2's domain
|
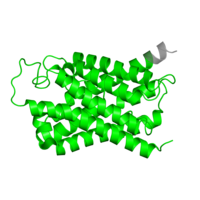
Figure 4: Structure of the MIP domain
|
The MIP domain is associated with aquaporins and other transporter proteins(4). The pore formed by the domain can be used for the transport of water or small neutral solutes. Greater divergence from human structure may be related to use of the homologs for transport of non-water substrates, such as has occurred in plants (5).
|
References
1. University of Verona.http://dp.univr.it/~laudanna/LCTST/styled-21/styled-26/. Acessed March 9th,2017.
2. The Pfam protein families database: towards a more sustainable future: R.D. Finn, P. Coggill, R.Y. Eberhardt, S.R. Eddy, J. Mistry, A.L. Mitchell, S.C. Potter, M. Punta, M. Qureshi, A. Sangrador-Vegas, G.A. Salazar, J. Tate, A. BatemanNucleic Acids Research (2016) Database Issue 44:D279-D285. <http://pfam.xfam.org/>
3. Letunic I, Doerks T, Bork P. SMART: recent updates, new developments and status in 2015.Nucleic Acids Res 2014; doi:10.1093/nar/gku949<http://smart.embl-heidelberg.de/>
4.Family:MIP. http://pfam.xfam.org/family/PF00230#tabview=tab0. Accessed March 9th,2017.
5. Maurel, C., Verdoucq, L., Luu, D. T., & Santoni, V. (2008). Plant aquaporins: Membrane channels with multiple integrated functions. Annual Review of Plant Biology, 59, 595-624. doi:10.1146/annurev.arplant.59.032607.092734
2. The Pfam protein families database: towards a more sustainable future: R.D. Finn, P. Coggill, R.Y. Eberhardt, S.R. Eddy, J. Mistry, A.L. Mitchell, S.C. Potter, M. Punta, M. Qureshi, A. Sangrador-Vegas, G.A. Salazar, J. Tate, A. BatemanNucleic Acids Research (2016) Database Issue 44:D279-D285. <http://pfam.xfam.org/>
3. Letunic I, Doerks T, Bork P. SMART: recent updates, new developments and status in 2015.Nucleic Acids Res 2014; doi:10.1093/nar/gku949<http://smart.embl-heidelberg.de/>
4.Family:MIP. http://pfam.xfam.org/family/PF00230#tabview=tab0. Accessed March 9th,2017.
5. Maurel, C., Verdoucq, L., Luu, D. T., & Santoni, V. (2008). Plant aquaporins: Membrane channels with multiple integrated functions. Annual Review of Plant Biology, 59, 595-624. doi:10.1146/annurev.arplant.59.032607.092734